Read our recent paper for more details: Saez et al., 2021
Many natural phages are unreliable and unfit for therapy. Obviously, only those phages that never eradicated their host can be found in nature today. When we compiled known S. aureus phages from public sources, we noticed that, while plaquing host ranges were generally large, the phages performed less well in other tests, such as suppressing bacterial growth in suspension or in biofilm assays.
A common problem is illustrated in the figure below: while phage 812 plaques similarly well on the 3 example strains, it suppresses growth in suspension only for the first strain. For the second strain, resistant outgrowth is detected after 9-12 hours, and the growth of the third one is only barely suppressed at all. Would that be a problem in the clinic? Shouldn’t the phages reduce the bacterial populations by many logs and keep them low to be effective therapeutics?
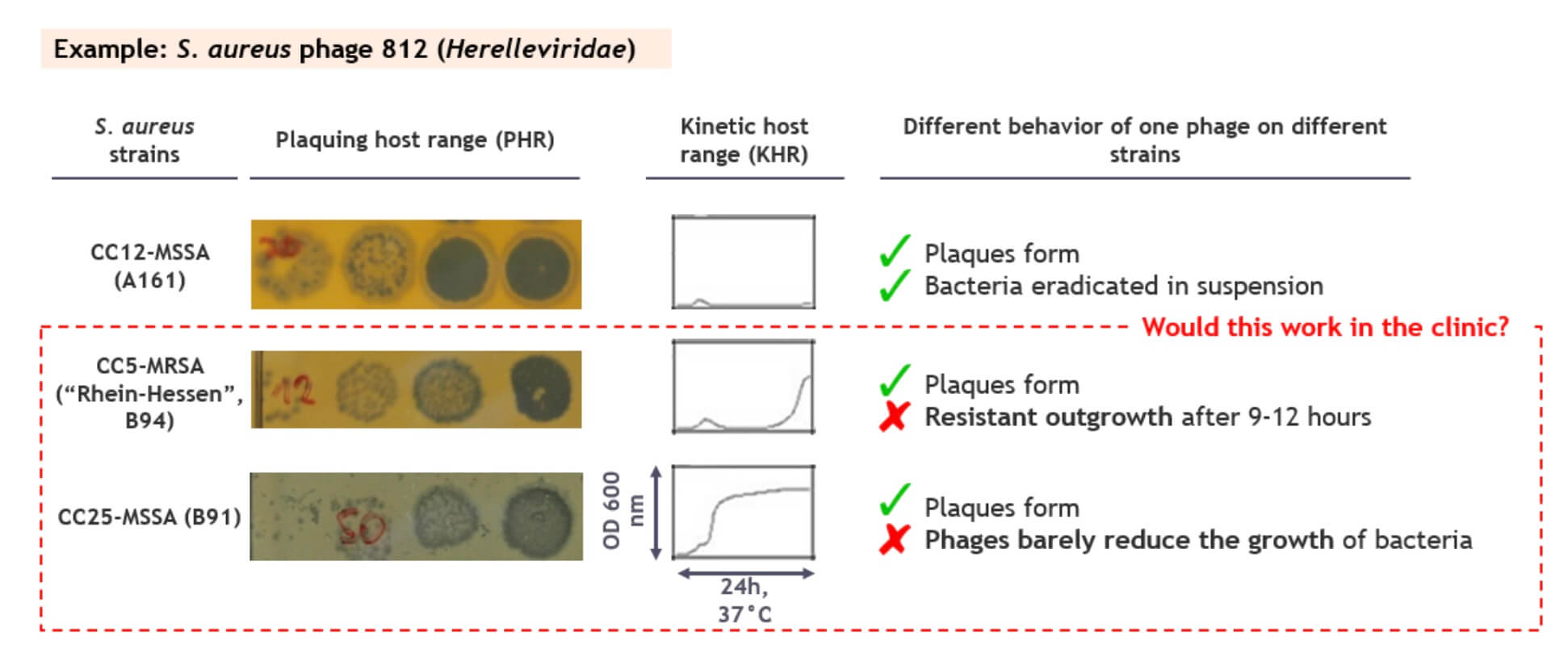
Figure 1. A high efficiency of plaquing does not always translate to high virulence, especially not for wild type phages. Image credit: PhagoMed Biopharma GmbH.
Therefore, we set out to improve the virulence, or lytic aggressiveness, of the phages, to close the gap between the plaquing host range and what we call the “kinetic host range”, KHR.
Phage breeding by homologous recombination upon superinfection (‘phage sex’)
Of the many published methods to improve phages over the wild types, we chose to let natural evolution go wild. We started with a method similar to the one called Appelman’s protocol described in the Burrowes 2019 paper. We changed many aspects — for example because the original Appelman’s paper from 1921 is optimized for titering, not for breeding. Also, we compiled a panel of S. aureus host strains with a highly controlled diversity, to direct the host range of the bred phages to as many human-relevant clonal complexes as possible.
To the best of our knowledge, the first scientist to systematically cross-breed phages by this method was Alfred Hershey, one of the founding fathers of molecular biology, in 1949. Hershey cross-bred E. coli phages by inducing superinfections and selecting for combined host range and plaque-type phenotypes — not unsimilar to what Mendel did with his peas.
We optimized the technique of inducing phage superinfections and combined it with efficient selection filters. This virtually doubled the kinetic host range of some phages compared to the best ancestor, essentially closing the gap between plaquing and kinetic host range.
ɛ2-phages specific to S. epidermidis were already used to help a patient
We also bred S. epidermidis phages on a diverse panel of S. epidermidis strains, largely from implant infections. James Doub of the University of Maryland asked if we had S. epidermidis phages for a patient with an infected implant, who could not undergo the (normally required) full prosthesis exchange. We tested wild type and ɛ2-phages against the patient strain, and the ɛ2-phages were clearly superior. The patient was treated and is under remission for more than 6 months – the first time ɛ2-phages were used to help a patient!
What did we learn from promoting phage sex?
-
A phagogram by plaquing may not be the appropriate (sole) predictor of phage efficacy in vivo.
-
Phage recombination upon superinfection seems to be a very frequent event, as long as the parental phages have a minimum degree of homology.
-
Counterintuitively, the host range expansion in our phages was not driven by changes in the tail spike proteins, which are often cited as the main drivers of host range.
-
As the genetic recombinations are random, no a priori knowledge of phage genes is required – in this respect, the method has huge advantages over more targeted genetic engineering approaches.
-
The breeding also enables an approach to patenting not possible for natural phages.
Concluding thoughts
Phages did not evolve to eradicate bacteria, but to propagate themselves. Just imagine applying the power of systematic cross-breeding with selection for biofilm eradication, avoidance of bacterial resistance, propagation in physiological environments, pharmacokinetics and tissue distribution, etc. We believe this is just the beginning, and that systematic phage cross-breeding combined with selection for useful pharmaceutical properties will hugely improve the prospects of phage therapy.
What’s next?
We are looking for collaborations to extend the application of ɛ2-phages. Reach out if you are interested!






